Scan workflow¶
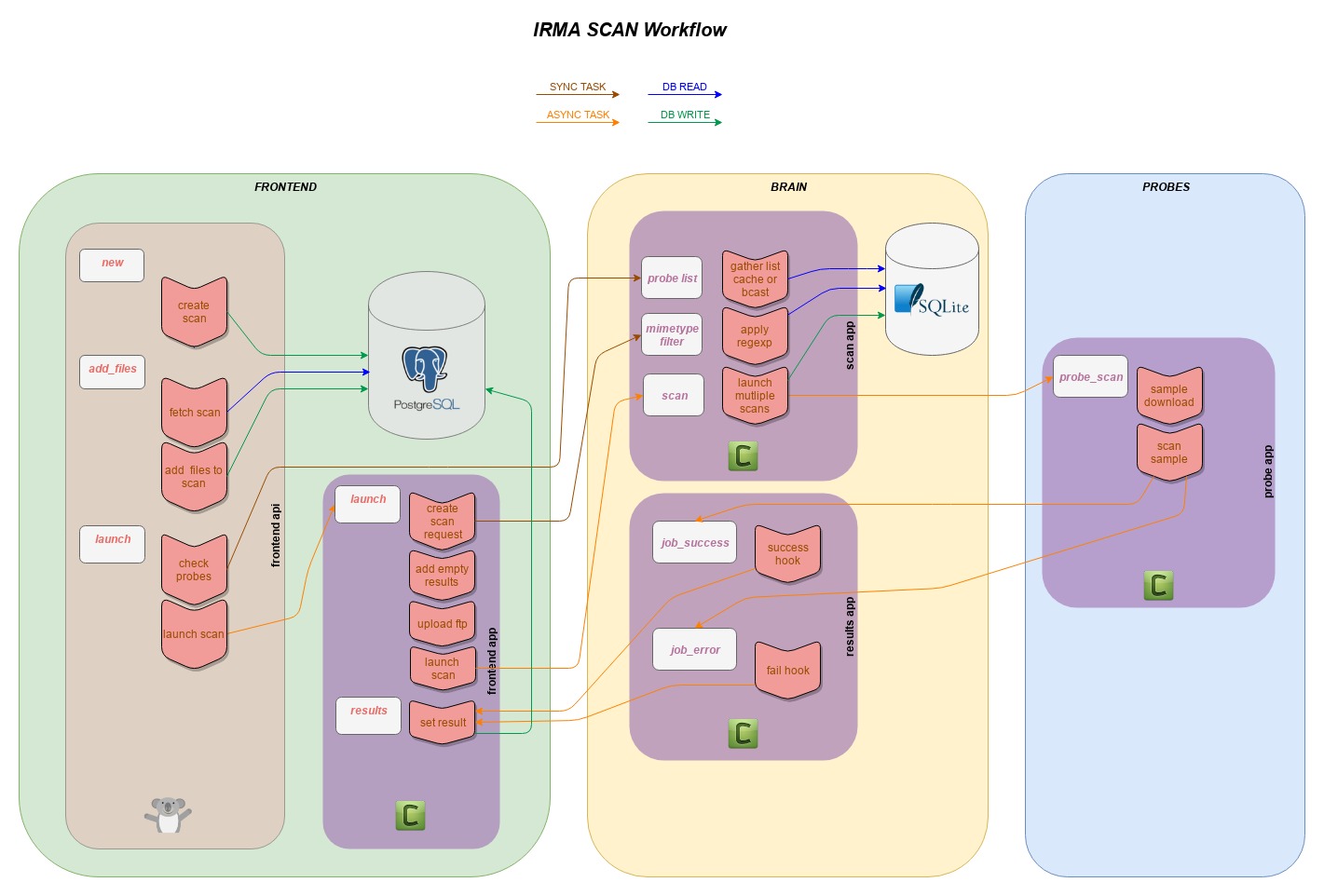
Frontend API Part (frontend_api/uwsgi+hug)¶
- A new scan object is created in PostgreSQL database.
- Files are uploaded to the WEB API, stored on Filesystem and registered in PostgreSQL database.
- Scan is launched, an asynchronous task is launched on Frontend celery.
Frontend Celery Part (frontend_app/celery)¶
- Used probes are filtered according to scan options (selected probes, mimetype filtering).
- Empty results are created in PostgreSQL database (one per probe per file).
- Each file is uploaded to SFTP server.
- For each file uploaded a scan task on Brain is launched with the file probelist (according to scan option force some results could already be present).
Brain Celery Part (scan_app/celery)¶
- A new scan object is created in SQLite database to track jobs (for canceling).
- Each file is send for analysis in sent for analysis in every probe selected (each time a probe is available in IRMA, it registers itself to the brain and open a RabbitMQ Queue named with its probe name, probe list is retrieved by listing active queues).
- Two callbacks are set on every probe scan tasks, one for success and the other for failure.
Probe Celery Part (probe_app/celery)¶
- Scan task is received with a file id.
- File is downloaded as temporary file.
- File is scanned by the probe.
- Results are sent back to Brain to one of the two callbacks set.
Brain Celery Part (result_app/celery)¶
- successful results are marked as completed in SQLite database.
- successful results are forwarded to Frontend.
- error are marked as completed in SQLite database.
- As there is no result, an error message is generated to tell the Frontend the particular job for the file and probe failed.
Frontend Celery Part (frontend_app/celery)¶
- Results is received for each file and probe.
- Results are updated in PostgreSQL database.
- If scan is finished, a scan flush task is launched on Brain to delete files on SFTP server.